Graph-Based Community Detection in Hepatitis C Patient Data Using KNN Graph Construction and the Louvain Algorithm
DOI:
https://doi.org/10.29304/jqcsm.2026.18.12452Keywords:
Hepatitis C Virus, Clustering Techniques, KNN Algorithm, Community Detection, Louvain AlgorithmAbstract
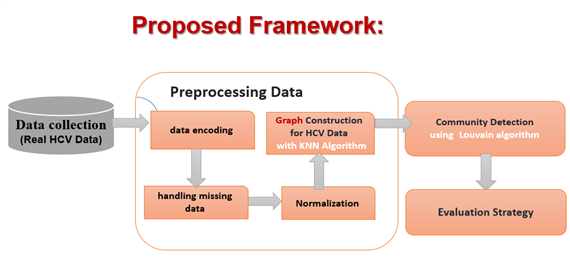
Analyzing medical data to identify clinically relevant patient groups remains a complex task, particularly for hepatitis C virus patients, as traditional clustering methods often struggle to capture heterogeneous patient relationships and may rely on specific geometric assumptions or be constrained by the number of groups to be found. The proposed framework for detecting patient communities combines local similarity representation and global structural analysis. This framework builds a network of patient similarity using the K-Nearest Neighbors (KNN) algorithm, followed by the detection of patient communities through the Louvain algorithm. The proposed method is implemented on two real hepatitis C virus (HCV) datasets: the Egyptian HCV dataset and the HCV (UCI) dataset. The results of the experiments show effective detection of coherent and stable patient communities. The highest modularity values reach 0.740 and 0.805 for the Egyptian and UCI datasets respectively, at a neighborhood value of K=3. In addition, in this context, low distortion values indicate strong cohesion within the detected communities. Overall, the results confirm that graph-based patient community discovery provides a powerful alternative to traditional clustering techniques for medical data analysis. The proposed framework enables reliable and stable discovery of clinically relevant and interpretable latent patient subgroups.
Downloads
References
M. A. A. Mohammed M. El Behery, Ahmed I. Elghwab, Ashraf A. Tabll, Elsherbeny H. Elsayed, “Review Article: Egyptian Efforts to Control Hepatitis C Virus in Egypt .,” 2025.
N. E. Izere Salomon, Sibomana Olivier, “Advancing Hepatitis C Elimination in Africa: Insights from Egypt.,” 2024.
M. B. Mojca Matičič, “Towards eliminating hepatitis C as a public health threat: different speeds, different needs,” 2024.
H. H. A. Elbahrawy, Ashraf, Marwa K. Ibrahim, Ahmed Eliwa, Ali Madian Mohamed Alboraie, “Current situation of viral hepatitis in Egypt,” 2021.
I. Gomaa, Asmaa, Mohamed Gomaa, Naglaa Allam, “Hepatitis C Elimination in Egypt: Story of Success.,” 2024. [Online]. Available: https://doi.org/10.3390/pathogens13080681
W. K. Al-hamoudi and Department, “Management of Hepatitis C Genotype 4 in the Liver Transplant Setting,” 2016. doi: 10.4103/1319-3767.182453.
S. Mahmud, Z. Al-Kanaani, H. Chemaitelly, K. Chaabna, S. P. Kouyoumjian, and L. J. Abu-Raddad, “Hepatitis C virus genotypes in the Middle East and North Africa: Distribution, diversity, and patterns,” 2018. doi: 10.1002/jmv.24921.
H. A. A. Khudhair* and K. R. H. , Ali A. H. Albakaa, “Detecting the Prevalence of Hepatitis C Virus among Iraqi People.,” 2023.
S. D. Warkad, K. Song, and D. Pal, “Developments in the HCV Screening Technologies Based on the Detection of Antigens and Antibodies,” 2019.
G. S. Vincenzo Moscato, “Community detection over feature-rich information networks: An eHealth case study,” ELSEVIER, 2022, [Online]. Available: https://doi.org/10.1016/j.is.2022.102092
C. Krantsevich, “Digital medicine and the curse of dimensionality,” pp. 1–8, 2021, doi: 10.1038/s41746-021-00521-5.
L. G. Buch, Amanda M., Conor Liston, “Simple and Scalable Algorithms for Cluster-Aware Precision Medicine,” 2023.
C. Simpson et al., “Lifting the curse from high-dimensional data : automated projection pursuit clustering for a variety of biological data modalities,” pp. 1–20, 2025, doi: 10.1093/gigascience/giaf052.
J. B. L. RYAN A. ROSSI, DI JIN, SUNGCHUL KIM, NESREEN K. AHMED, DANAI KOUTRA, “On Proximity and Structural Role-based Embeddings in Networks: Misconceptions, Techniques, and Applications.,” 2020.
L. Brahim, M. Mouad, C. Chihab-eddine, and I. Ali, “A SURVEY ON COMMUNITY DETECTION : APPLICATIONS , ALGORITHMS , AND CHALLENGES,” vol. 102, no. 12, pp. 4923–4945, 2024.
I. K. Isa Inuwa-Dutse, Mark Liptrott, “A multilevel clustering technique for community detection.,” 2021.
W. Zhao, J. Luo, T. Fan, and Y. X. Ren, Yan, “Analyzing and visualizing scientific research collaboration network with core node evaluation and community detection based on network embedding,” 2021, [Online]. Available: https://doi.org/10.1016/j.patrec.2021.01.007
2 MEHRDAD ROSTAMI 1, MOURAD OUSSALAH 1, 2, (Senior Member, IEEE), KAMAL BERAHMAND 3, AND VAHID FARRAHI 1, “Community Detection Algorithms in Healthcare Applications: A Systematic Review.,” 2023, IEEE.
J.-K. H. Goudet, Olivier, B´ eatrice Duval, “Population-based Gradient Descent Weight Learning for Graph Coloring Problems,” 2020.
J. Ren, X. Lyu, J. Guo, X. Shi, Y. Zhou, and Q. Li, “CDSKNN XMBD : a novel clustering framework for large ‑ scale single ‑ cell data based on a stable graph structure,” J. Transl. Med., pp. 1–11, 2024, doi: 10.1186/s12967-024-05009-w.
V. D. Blondel, J. Guillaume, and E. Lefebvre, “Fast unfolding of communities in large networks,” pp. 1–12, 2008.
R. G. Dipika Singh, “NI-Louvain: A novel algorithm to detect overlapping communities with influence analysis.pdf,” 2022, Elsevier.
S. Seth, S. Mallik, T. Bhadra, and Z. Zhao, “Dimensionality Reduction and Louvain Agglomerative Hierarchical Clustering for Cluster-Speci fi ed Frequent Biomarker Discovery in Single-Cell Sequencing Data,” vol. 13, no. February, pp. 1–17, 2022, doi: 10.3389/fgene.2022.828479.
W. Makelaardij, “A cost-based multi-layer network approach for the discovery of patient phenotypes Clara,” 2023.
A. M. Elshewey et al., “Optimizing HCV Disease Prediction in Egypt : The hyOPTGB Framework,” 2023.
H. M. Farghaly, M. Y. Shams, and T. A. El-hafeez, “Hepatitis C Virus prediction based on machine learning framework : a real-world case study in Egypt,” Knowl. Inf. Syst., vol. 65, no. 6, pp. 2595–2617, 2023, doi: 10.1007/s10115-023-01851-4.
B. Pankratz and P. Prałat, “Performance of community detection algorithms supported by node embeddings,” 2024.
A. A. Permana and R. M. Yaputra, “Analyzing breast cancer comorbidities : a network approach using community detection algorithms,” Appl. Netw. Sci., 2024, doi: 10.1007/s41109-024-00644-0.
J. Bambi et al., “Approaches to Extracting Patterns of Service Utilization for Patients with Complex Conditions : Graph Community Detection vs . Natural Language Processing Clustering,” no. 1, pp. 1884–1900, 2024.
M. Bampa, I. Miliou, B. Jovanovic, and P. Papapetrou, “M-ClustEHR : A multimodal clustering approach for electronic health records,” vol. 154, no. May, 2024.
A. Sharma, T. Khade, and S. M. Satapathy, “A cross dataset meta-model for hepatitis C detection using multi- dimensional pre-clustering,” pp. 1–17, 2025.
N. J. van E. Traag, V. A., L. Waltman, “From Louvain to Leiden: guaranteeing well-connected communities,” 2019.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2026 Bashair Mohammed Obaid, Hayder K. Fatlawi

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.